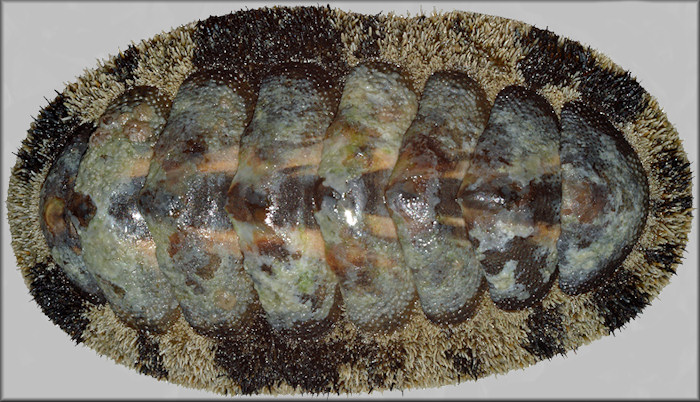
Congratulations to Ph.D. candidate Rebecca Varney for winning a best talk award at the World Congress of Malacology for her presentation on the genome of the chiton Acanthopleura granulata (Mollusca: Polyplacophora)!
Abstract of the talk: Molluscan genomic resources continue to increase, but with a marked bias toward bivalves and gastropods. Chitons are models for molluscan biomineralization, producing shell valves, sclerites, and radulas with teeth coated with magnetite (iron). Here we present the first genome of a chiton, the first from clade Aculifera. A single individual of Acanthopleura granulatawas collected from the Florida Keys and DNA was extracted from foot tissue using a phenol-chloroform-based approach optimized to provide clean, high-molecular weight DNA, with modifications that proved critical for successful nanopore sequencing. A highly cost-effective hybrid assembly strategy was employed combining Illumina short reads and Oxford Nanopore long reads in MaSuRCA followed by Bionano SAPHYR optical mapping for scaffolding. Afterreduction of heterozygosity with Redundans, removal of bacterial scaffolds with BlobTools, and polishing of the final assembly with 4 rounds of PILON, we produced a 605.9 Mbp assembly in 73 scaffolds with an N50 of 23.9Mbp and a BUSCO completeness score of 96.2%. Annotation was carried out with multiple strategies compared. Many (but not all) genes hypothesized to be part of the conchiferan mollusc biomineralization toolkit are also present in chitons, shedding light on the possible ancestral molluscan suite of biomineralization genes. Additionally, analysis of known iron-associated proteins determined some sequence variation in known iron-binding proteins in chitons. Future screening for iron-regulated domains in the genome will be combined with 3’-biased sequencing of transcriptomes from the developing chiton radula to determine the gene expression patterns underlying magnetite mineralization.